Introduction
This is my bespoke pipeline for processing MinION metabarcoding data, focused specifically on ITS2 in vascular plants. It hasn’t yet been reviewed by anyone with any special expertise in this area, but you may nevertheless find it useful.
I find the documentation of bioinformatics tools and pipelines generally lacking, or at least targeted at people with more background knowledge than I have. To compensate for this, I move very slowly through a protocol, and examine intermediate outputs with standard text-processing tools. This is essential for my learning process, but may be tedious if you do have the understanding the bioinformaticians expect you to have.
In any case, what follows is a very detailed walk through of my pipeline. Now that it works I’ll convert it to a single script to run through all the bits below.
Library Preparation
The wet lab protocol we’re using is very similar to that used by ONTBarcoder (Srivathsan et al., 2024), except we’re using ITS2 instead of COI. The ONTBarcoder pipeline isn’t limited to COI, but it does expect the target amplicon to be a consistent length. That’s not the case with ITS2, so I put together the following pipeline.
Our wet lab protocol consists of:
- DNA extraction
- amplification with ITS2-S2F and ITS4, both primers with a 9bp index
- PCR products further processed with the Nanopore ligation kit SQK-LSK114
The completed library is:
LSK adapter - Forward Index - ITS2-S2F - amplicon -
- ITS4 - Reverse Index - LSK adapterWe expect the length of individual products to be 218-378 bp. This accounts for documented variation in IT2 length (160-320bp) plus primers and adapters. Adding a bit of buffer, I’ll consider sequences in the range 160-500bp as a potentially ‘valid’ read.
The LSK Adapter sequence is:
TTTTTTTTCCTGTACTTCGTTCAGTTACGTATTGCT
GGACATGAAGCAAGTCAATGCATAACGPrimers:
- ITS-S2F: ATGCGATACTTGGTGTGAAT
- ITS4: TCCTCCGCTTATTGATATGC
Our indexes are 9 bp long:
| Forward | Reverse |
|---|---|
| GGAGAAGAA | CTCACAAGG |
| CAGAGAGAA | AAGAGCAGG |
| ACTCCAGAA | TAACTGCGA |
| TCCAAGGAA | GTCATCCGA |
| … | … |
Dorado will trim the LSK adapters, either as part of basecalling (which is the default) or in a separate step. However, in my early trials this seems to leave an inconsistent residue at the ends of the reads which was (initially at least) kind of confusing. (this turned out to be a non-issue with the approach I took).
Another issue is a small proportion of reads are concatenated into chimeras during sequencing. i.e., two sequences are read as one long one, containing two sets of amplicons along with their flanking primers and indexes. ONTBarcoder deals with this by splitting long sequences into two, and then treating each half as a separate read. In our first run only 1.1 percent of reads were long enough to be chimeras, so I’m just going to filter out these long reads and get on with my life.
With all that in mind, our pipeline will be:
- basecalling via Dorado
- extracting amplicons + flanking primers/indices via seqkit amplicon
- length filtering via seqkit seq
- demultiplex via demultiplex demux
- trim primers/indices via seqkit amplicon
- dereplicate via vsearch –fastx-uniques
- match to reference via vsearch –usearch-global
- further processing in R
Basecalling
In the end, it doesn’t really matter if we trim the adapters or not. We’re going to excise the amplicons in the next step, which will remove everything external to the indices.
In addition to the Dorado program, we need the basecalling models. We can download them with Dorado:
dorado download --model allThis downloads a whole load of models. The one we want will be in a
directory named dna_r10.4.1_e8.2_400bps_sup@v5.2.0. The particulars of
the model are encoded in the directory name:
- dna
- DNA sequence
- r10.4.1
- FLO-MIN114 flow cell
- e8.2
- chemistry type, Kit 14
- 400bps
- translocation speed
- sup
- super accurate
- v5.2.0
- model version
We will always use dna_r10.4.1_e8.2_400bps_sup, as these parameters are
set by the hardware we’re using. We may want to update to the newer model
versions as they are released.
dorado basecaller \
./path/to/model/dna_r10.4.1_e8.2_400bps_sup@v5.2.0 \
./path/to/pod5 \
--device cuda:all --emit-fastq --no-trim \
> output_dir/basecalls.fastqNB: --device cuda:all for specifying a GPU.
This takes about 20 minutes on the cluster (all times will depend on the size of your read files of course; I include mine here to give you an idea of the relative time required at each step).
Check the number of reads:
wc -l < output_dir/basecalls.fastq 4362908Remember, four lines per read, so total reads = 1,090,727.
Amplicon Selection
In initial trials, this worked best if I select the 9 bp external to each primer. i.e., exactly matching the expected size of the barcodes. This means we’ll drop any reads where there was an indel in the index. Which is probably a good thing to do in any case.
seqkit amplicon -F ATGCGATACTTGGTGTGAAT \
-R TCCTCCGCTTATTGATATGC -r -9:9 -f \
output_dir/basecalls.fastq -m 4 > \
output_dir/amplicons_9m4.fastq-Fand-R- primer sequences
-r -9:9 -f- extract the sequence starting 9 bp before the forward primer and extending 9 bp after the reverse primer
-m 4- allow a maximum of four mismatches in primer sites
Takes about 2 minutes.
As a quick sanity check, look at the first read in the output:
head -2 amplicons_9m4.fastq@d2b8520e-173a-441d-ab34-b9f8ea53d679 qs:f:9.27552 [ ... ]
TTCCAACAAATGCGATACTTGGTGTGAATTGCAGAATCCCGTGAACCATCGAGTCTTTGAACGCAAGTTGCGCCCGAGGCCTCCTGGTCGAGGGCACGTCTGCCTGGGTGTCACGCATCGTCGCCCCCACTCCCCTCGGCTCACGAGGGCGGGGGCGGATACTGGTCTCCCGCGCGCTCCCGCCCGTGGTTGGCCTAAAATCGAGTCCTCGGCGACGGTCGCCACGACAAGCGGTGGTTGAGATAGCTCGATGGTCGGTGTGTGTCGTTGCCGCCCTGGGGAACTCCCGGTACCGCCGAGCATTTGGCTTCAGGGATGCTCGCTGGGACCCCAGCGTGGCAGGACCCTCTAAGCATATCAATAAGCGGAGGATGTCGTGTTAnd compare it to the raw read:
grep -A 1 @d2b8520e basecalls_2026.fastq@d2b8520e-173a-441d-ab34-b9f8ea53d679 qs:f:9.27552 [ ... ]
> TTAAGTTGTAACCTACTCGACTTTCAGTTACGTATTGCTAACACGACATCCTCCGCTTATTGATATGCTTAGAGGGTCCTGCCACGCTGGGGTCCCAGCGAGCATCCCTGAAGCCAAATGCTCGGCGGTACCGGGAGTTCCCCAGGGCGGCAACGACACACACCGACCATCGAGCTATCTCAACCACCGCTTGTCGTGGCGACCGTCGCCGAGGACTCGATTTTAGGCCAACCACGGGCGGGAGCGCGCGGGAGACCAGTATCCGCCCCCGCCCTCGTGAGCCGAGGGGAGTGGGGGCGACGATGCGTGACACCCAGGCAGACGTGCCCTCGACCAGGAGGCCTCGGGCGCAACTTGCGTTCAAAGACTCGATGGTTCACGGGATTCTGCAATTCACACCAAGTATCGCATTTGTTGGAAAGCAATGCGTTI visually matched the raw read (reverse complimented in this case, the top
line) to the output from amplicon (the middle line):
AACGCATTGCTTTCCAACAA ATGCGATACTTGGTGTGAAT TGCAGAATCC [ ... ]
TTCCAACAA ATGCGATACTTGGTGTGAAT TGCAGAATCC [ ... ]
INDEX Forward Primer AMPLICON
[ ... ] GTGGCAGGACCCTCTAA GCATATCAATAAGCGGAGGA TGTCGTGTT AGCAATACG [ ... ]
[ ... ] GTGGCAGGACCCTCTAA GCATATCAATAAGCGGAGGA TGTCGTGTT
[ ... ] AMPLICON Reverse Primer (rc) INDEXAll is as expected, so we can proceed with the analysis.
Length Filtering
Extract read lengths:
awk 'NR % 4 == 2 { print length }' \
output_dir/amplicons_9m4.fastq > \
output_dir/9m4_lengths.txtReview read lengths in R:
reads <- read.table("output_dir/9m4_lengths.txt",
sep = "\t")
readsTrim <- reads[reads <= 1000, ]
hist(readsTrim, breaks = 50, xlab = "Read Length",
ylab = "Number of Reads", main = "")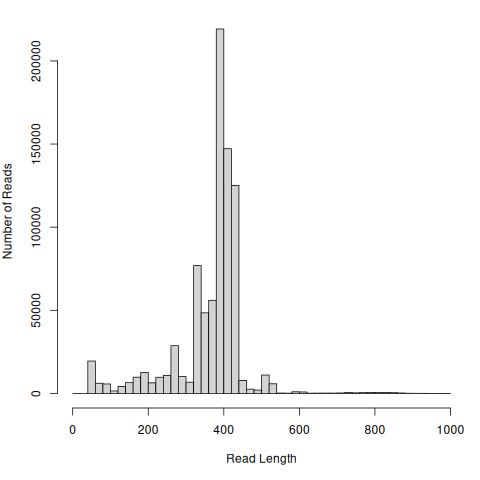
Note in this case I’m not plotting reads longer than 1000 bp, of which there are 160 (from 848,255 total, less than 0.01%).
Filtering my reads to 200-500 bp drops ~10% of the total:
seqkit seq -m 200 -M 500 output_dir/amplicons_9m4.fastq > \
output_dir/amp9M4_500.fastqwc -l < amp9M4_500.fastq3031992757,998 reads retained. Takes 10 seconds.
Demultiplexing
Note: demultiplex appends output to existing files. So we need to remove
any files from a previous run before we re-run it on the same files (e.g.,
when tweaking parameters).
for_indexes.tsv:
F1 GGAGAAGAA
F2 CAGAGAGAA
F3 ACTCCAGAA
F4 TCCAAGGAA
...rev_indexes.tsv:
R1 GAGTACGGA
R2 AGAACCGGA
R3 TAACTGCGA
R4 GTCATCCGA
...rm -r output_dir/demux9/*
demultiplex demux -p output_dir/demux9 -s 1 -e 9 \
for_indexes.tsv output_dir/amp9M4_500.fastq-s 1 -e 9- match indexes against base pairs [S]tarting at bp 1 and [E]nding at bp 9.
Takes one minute.
I’m not sure how to tell demultiplex to look in the last 9 bp for the
barcodes. So I’m going to reverse compliment the sequences here. This is
very quick and can be done on the head node:
for FILE in output_dir/demux9/*fastq
do
OUT=$(basename $FILE .fastq)_rc.fastq
seqkit seq -r -p -t dna $FILE > output_dir/demux9/$OUT
doneNow we have the sequences in the right order, proceed to demultiplex a second time with the reverse index:
rm output_dir/demux9/demux9b/*
for FILE in output_dir/demux9/*_rc.fastq
do
demultiplex demux -p output_dir/demux9/demux9b \
-s 1 -e 9 rev_indexes.tsv $FILE
done
Takes one minute.
Checking one sample:
wc -l < output_dir/demux9/demux9b/amp9M4_500_F8_rc_R11.fastq 105244i.e., 26,311 reads for this sample.
Trimming
Return to seqkit amplicon to trim our primer sites:
rm output_dir/trim/*
for FILE in output_dir/demux9/demux9b/*_rc_R*
do
OUT=$(basename \$FILE .fastq)
OUT=${OUT/_rc_/_}_trim.fastq
seqkit amplicon -F ATGCGATACTTGGTGTGAAT \
-R TCCTCCGCTTATTGATATGC \
-r 20:-20 $FILE -m 4 > output_dir/trim/${OUT}
done-r 20:-20- Extract the first and last 20 base pairs from each amplicon (i.e., the primer sites)
-m 4- maximum four mismatches in primer sites
Dereplication
We are expecting a small number of distinct sequences, each of which may be present in very high numbers. We don’t want to match each of the duplicate sequences to the database, so the next step is to extract a single representative of each unique sequence.
for FILE in output_dir/trim/*trim.fastq
do
NAME=$(basename $FILE)
OUT=${NAME%trim.fastq}
vsearch --fastx_uniques $FILE \
--sizeout --quiet --minuniquesize 2 \
--fastqout output_dir/derep/${OUT}dr.fq
done
--fastqout-
output
fastqformats --sizeout- include number of replicates in output
--minuniquesize 2- Discard singleton sequences, as they are likely to be sequencing errors
Takes a few seconds.
Note that as we’ve requested --sizeout, every retained read will have the
string size= in its header line. Counting these tells us how many unique
sequences were found in a sample.
grep -c "size=" output_dir/amp9M4_500_F8_R11_dr.fq683Summing the values listed for size tells us how many of the original
reads we retained (i.e., after excluding singletons).
awk 'BEGIN { FS = "size=" }
/size/ { tot = tot + $2}
END { print tot}' output_dir/amp9M4_500_F8_R11_dr.fq6542Recall we had 26,311 after dereplication, so 20k reads were singletons.
Taxon Matching
globaldb.fasta is from (Quaresma et al., 2024). It’s comprehensive, based on
a lightly cleaned set of sequences from GenBank. A locally curated,
validated database will be better whenever available. If it’s not, be
sure to examine the individual reference sequences matched.
There are quite a few output options. We can get everything we need from
--biomout or --uc.
for FILE in ${output_dir}/derep/*dr.fq
do
NAME=$(basename $FILE)
OUT=$(basename $NAME _dr.fq)
vsearch --usearch_global ${output_dir}/derep/$NAME \
--db globaldb.fasta \
--uc ${outout_dir}/taxon/$OUT.uc \
--biomout ${output_dir}/taxon/$OUT.biom \
--maxaccepts 100 --id 0.98 --sizeout
done--maxaccepts 100- by default, the search stops when the first target is found that passes the matching criterion (i.e., > 0.98 identity). This may not be the best match! Set to 0 to search the entire database, or another number N to force the search to consider at least the first N references that pass the criterion before deciding which one is the best.
--id 0.98- reject a sequence match if the pairwise identify is less than the 0.98
--sizeout- add abundance annotations to –uc file
Takes about an hour.
We can check the --uc output from the command line. Each query sequence
has a line in the file. The first column indicates Hits (matches) or No
matches. The ninth column includes the query sequence ID and the number of
replicates (i.e., size=). The 10th column indicates the taxon the query
matched to, in case of matches.
Unique query sequences:
wc -l < output_dir/amp9M4_500_F8_R11.uc683Matching queries:
grep -c '^H' output_dir/amp9M4_500_F8_R11.uc330Total sequences matched:
grep '^H' output_dir/amp9M4_500_F8_R11.uc | cut -f9 | \
awk 'BEGIN { FS = "size=" }
{ tot = tot + $2}
END { print tot}'2405Our reference has multiple sequences for many taxa. We can sum those up by species:
grep '^H' output_dir/amp9M4_500_F8_R11.uc | cut -f10 | \
sed 's/^.*s://' | sort | uniq -c | sort -h 128 Cucumis_sativus;
83 Trichophorum_pumilum;
53 Rhododendron_tomentosum;
39 Capsicum_frutescens;
15 Brassica_napus;
9 Solanum_lycopersicum;
2 Rhododendron_groenlandicum;
1 Rhododendron_ferrugineum;At this point the analysis moves from the realm of bioinformatics and back into the realm of botany. Further assessment is easiest (for me) to do in R. I’ll leave that for another post.